Hello all,
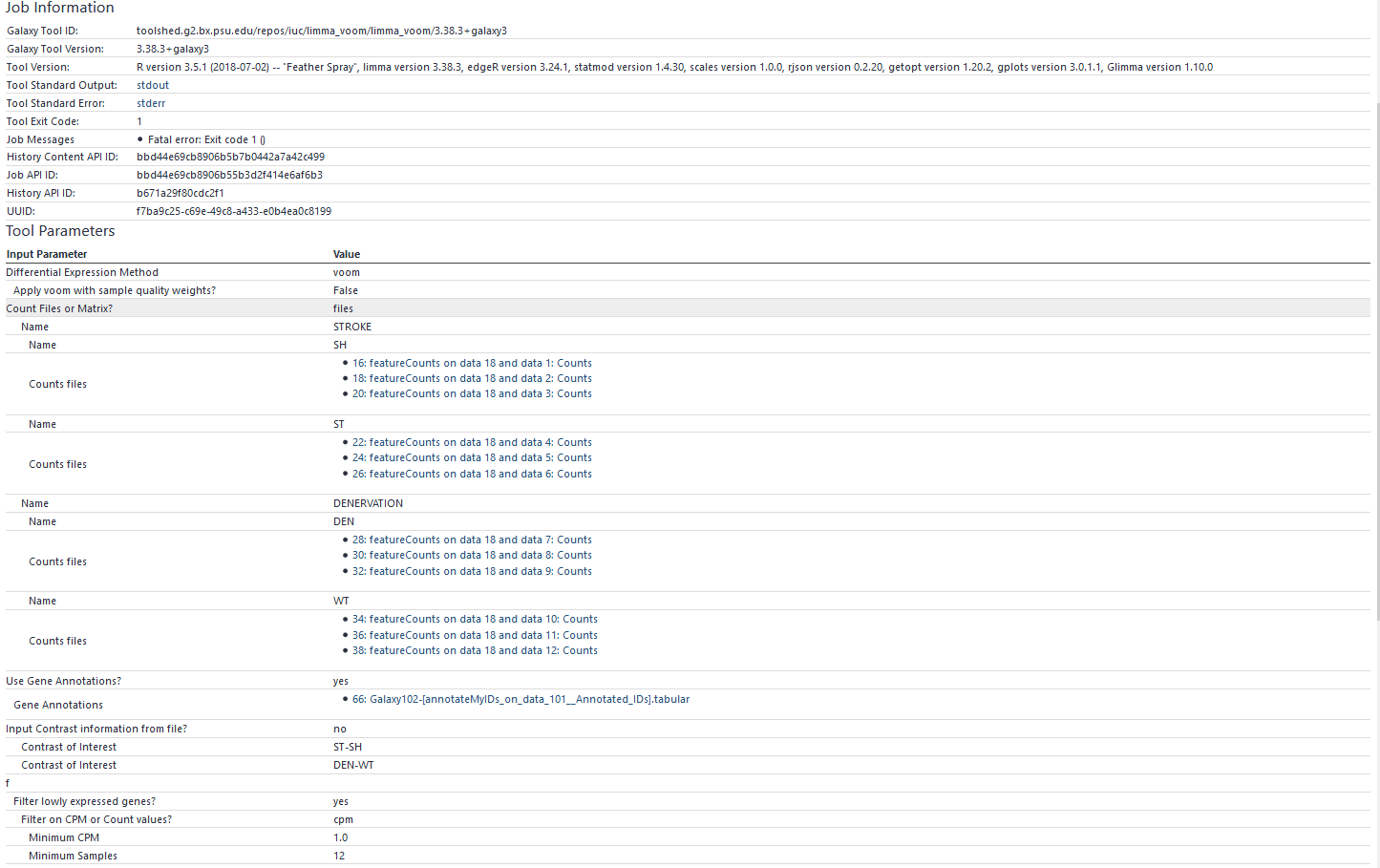
I’ve been struggling here for days now on attempting to get multiple factors to work in Galaxy for either EdgeR OR limma. I’ve searched every post about this and each says it’s a header issue. It is NOT a header issue. If I have only one factor and input the 5 separate groups within that one factor, each with it’s own count file it works fine and I can compare between each via contrasts. The count files have headers as GeneID for column1 and the unique sample name in column 2. However, when I attempt to add a separate Factor and include use the same files that worked in the previous step…I always get a duplicate row name error. I’ve tried even doing this via a count matrix file with all of the samples and counts…as well as a Factor info file; for both I followed the EXACT format as described on the package help section displayed if you scroll down. This did not work either.
Currently, I have absolutely no way to utilize the platform to peform DE analysis with the appropriate factor levels. Again, this is not a header issue with the counts files. I’ve tried this with both htseq counts (which I manually entered the header) AND featurecounts. These work fine if I only have one Factor level…as soon as I add an additional factor level it fails, despite being the exact same files that worked previously. I REALLY need help. I’d be greatly appreciative as my advisor needs this analysis for a grant application.
@Peter_Jon can you share your history. I will have a look, this sounds super strange. Btw. is deseq2 working?
I’ve only really tried to do this extensively with EdgeR and limma, but it didn’t work for DESeq2 either. How can I share my history to you?
“Error in .rowNamesDF<-(x, value = value) :
duplicate ‘row.names’ are not allowed
Calls: rownames<- … row.names<- -> row.names<-.data.frame -> .rowNamesDF<-
In addition: Warning message:
non-unique values when setting ‘row.names’:
Execution halted”
I could show you that all of my counts files have the correct format based upon the guide.
I just glanced over it as it is late here, but your files in both factors needs to be the same. Please consider the example of treated vs. un-treated and single vs. paired in this tutorial as example: https://training.galaxyproject.org/training-material/topics/transcriptomics/tutorials/ref-based/tutorial.html#analysis-of-the-differential-gene-expression for a 2 factor study.
I hope that helps,
Bjoern
This worked for both limma!, but NOT EdgeR unfortunately. And for DESeq2…I just can’t get to work in this way, no matter the combination. I tried primary factor (condition) = level 1 - ST2, ST3, ST4 level 2- SH1, SH2, SH3 level 3 - DEN1, DEN2, DEN3 level 4 - WT1, WT2, WT3 … then secondary factor (group) = level 1 - ST2, ST3, ST4, SH1, SH2, SH3 level 2 - DEN1, DEN2, DEN3. Gives me a fatal error saying
“Error in checkFullRank(modelMatrix) :
the model matrix is not full rank, so the model cannot be fit as specified.
One or more variables or interaction terms in the design formula are linear
combinations of the others and must be removed.”
After this, I attempted to run it so that the two separate Factors had no overlap and that also failed. Not sure what to do for DESeq2. Thank you for your help so far!