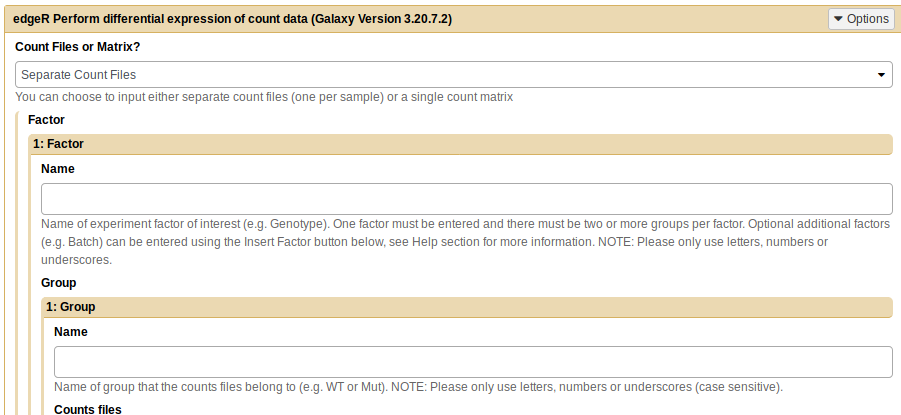
I am trying to use edgeR to perform a differential expression analysis. This is a paired analysis (samples are in pairs - their healthy vs their diseased tissues).
Galaxy Tool ID: toolshed.g2.bx.psu.edu/repos/iuc/edger/edger/3.20.7.2
Galaxy Tool Version: 3.20.7.2
Tool Version: R version 3.4.1 (2017-06-30) – “Single Candle”, edgeR version 3.20.7, limma version 3.34.9, scales version 0.5.0, rjson version 0.2.15, getopt version 1.20.0
I have 2 input files. The first is input-counts.txt and looks like so:
GeneID M1 M4 M7 M10 M13 N1 N4 N7 N10 N13
gene1 0 0 0 0 4 0 0 1 0 0
gene2 589 602 646 403 390 204 357 511 266 387
gene3 5 5 7 4 8 0 2 13 2 5
gene5 1 1 0 0 0 0 0 0 0 0
gene6 0 0 0 0 0 0 0 0 0 0
gene7 0 0 0 0 0 0 0 0 0 0
etc
My factor file looks as such:
Sample Patient Tissue
1 M1 1 M
2 M4 4 M
3 M7 7 M
4 M10 10 M
5 M13 13 M
6 N1 1 N
7 N4 4 N
8 N7 7 N
9 N10 10 N
10 N13 13 N
The aim of my analysis is to detect genes differentially
expressed between affected (M) and normal skin (N), adjusting for any differences between the patients.
In the edgeR window in Galaxy, in “Contrast” of interest", I have written: M-N.
However, edgeR doesn’t run and I get the following error:
Fatal error: Exit code 1 () Error in makeContrasts(contrasts = contrastData, levels = design) : The levels must by syntactically valid names in R, see help(make.names). Non-valid names:
I am confused as to what to do. I made this file in R. I have tried changing the levels() in R and I have tried putting “Tissue” in the second column but I still get the error each time.
Any help would be deeply appreciated.