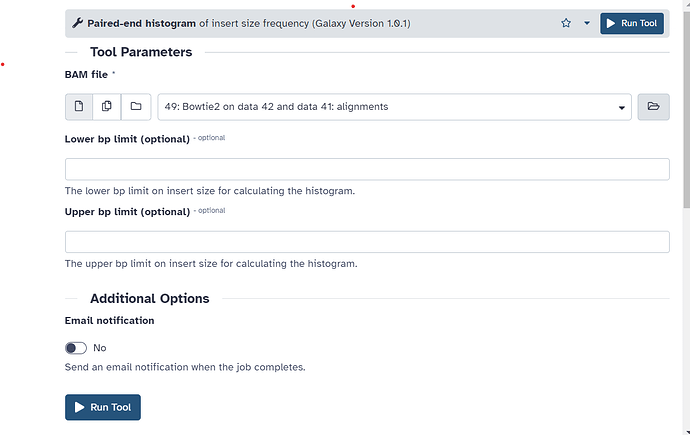
I have a project due for class and keep getting an error at the paired-end histogram step. I have asked the professor but he does not know what is wrong or how to help. Any ideas of what the error means and how I can fix it ?
This is the instructions
- Mapp the cutadapt output to the reference genome using Bowtie2
- The genome and gene annotations are in the “Sp5.0 Mapping Test” History that I already shared with you, and also posted in the shared DropBox folder IGV_Sp5_Genome_links.txt
- If you mapped to the whole genome, great! Otherwise, Map the reads to Sp5.0_Chromosome_NW_022145615.1.fa as if it was the entire genome. The chromosome and its corresponding index are also located in the DropBox folder.
- Filter duplicate reads with MarkDuplicates
- Visualize and analyze the distribution of read lengths with the tool Paired-end histogram****Peak call and visualization of normalized data
This is the error
An error occurred with this dataset:
format tabular database papHam1
Picked up _JAVA_OPTIONS: -Djava.io.tmpdir=/corral4/main/jobs/054/233/54233597/tmp -Xmx2488m -Xms256m Exception in thread “main” htsjdk.samtools.SAMFormatException: SAM validation error: ERROR: Record 941707, Read name LH00401:11:GW231120000:5:1105:14959: